ParSeq¶
- GitHub:
- PyPI:
- License:
- Author:
Konstantin Klementiev (MAX IV Laboratory)
Package ParSeq is a python software library for Parallel execution of Sequential data analysis. It implements a general analysis framework that consists of transformation nodes – intermediate stops along the data pipeline for data visualization, cross-data operations (e.g. taking average), providing user input and displaying status – and transformations that connect the nodes.
It provides an adjustable data tree model (supports grouping, renaming, moving and drag-and-drop arrangement), tunable data format definitions, plotters for 1D, 2D and 3D data, cross-data analysis routines and flexible widget work space suitable for single- and multi-screen computers. It also defines a structure to implement particular analysis pipelines as lightweight Python packages.
ParSeq is intended for synchrotron based techniques, first of all spectroscopy.
A screenshot of ParSeq-XAS (an EXAFS analysis pipeline) as an application example:
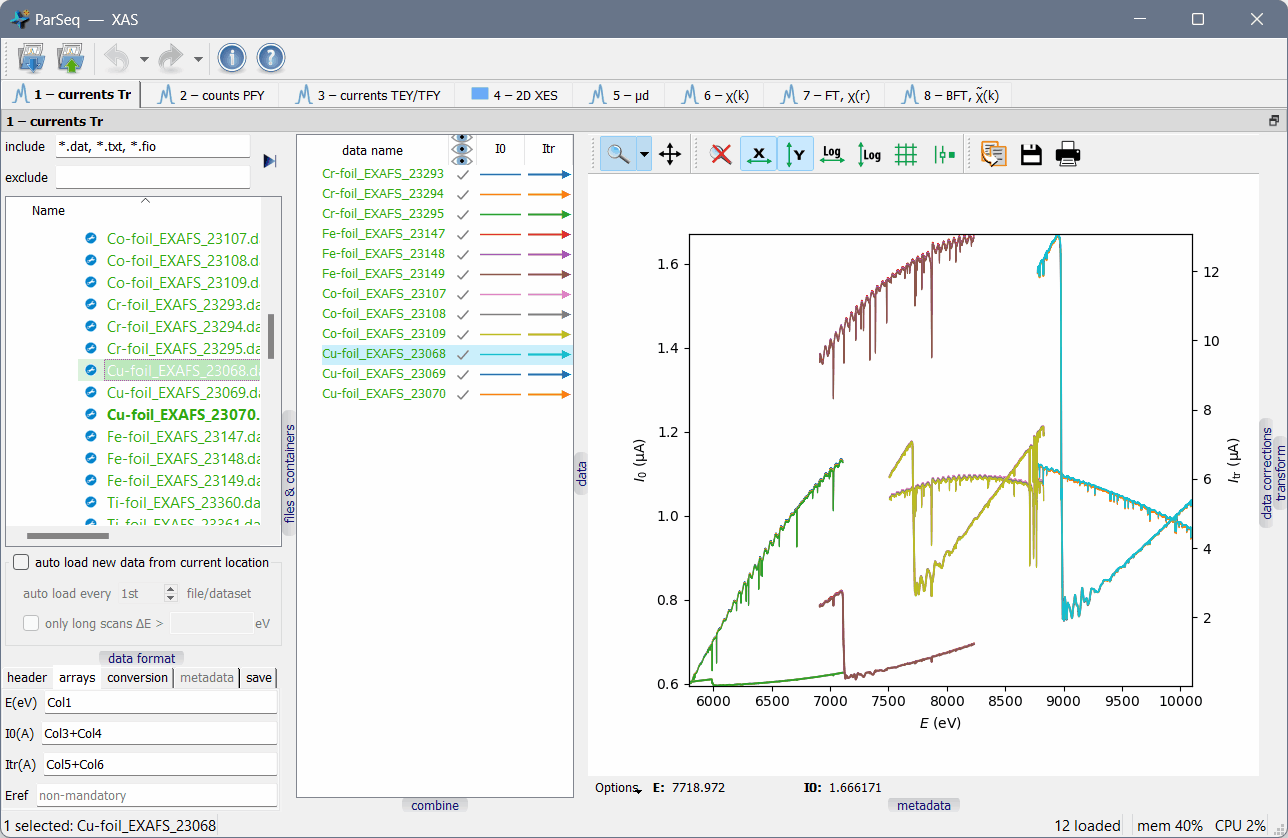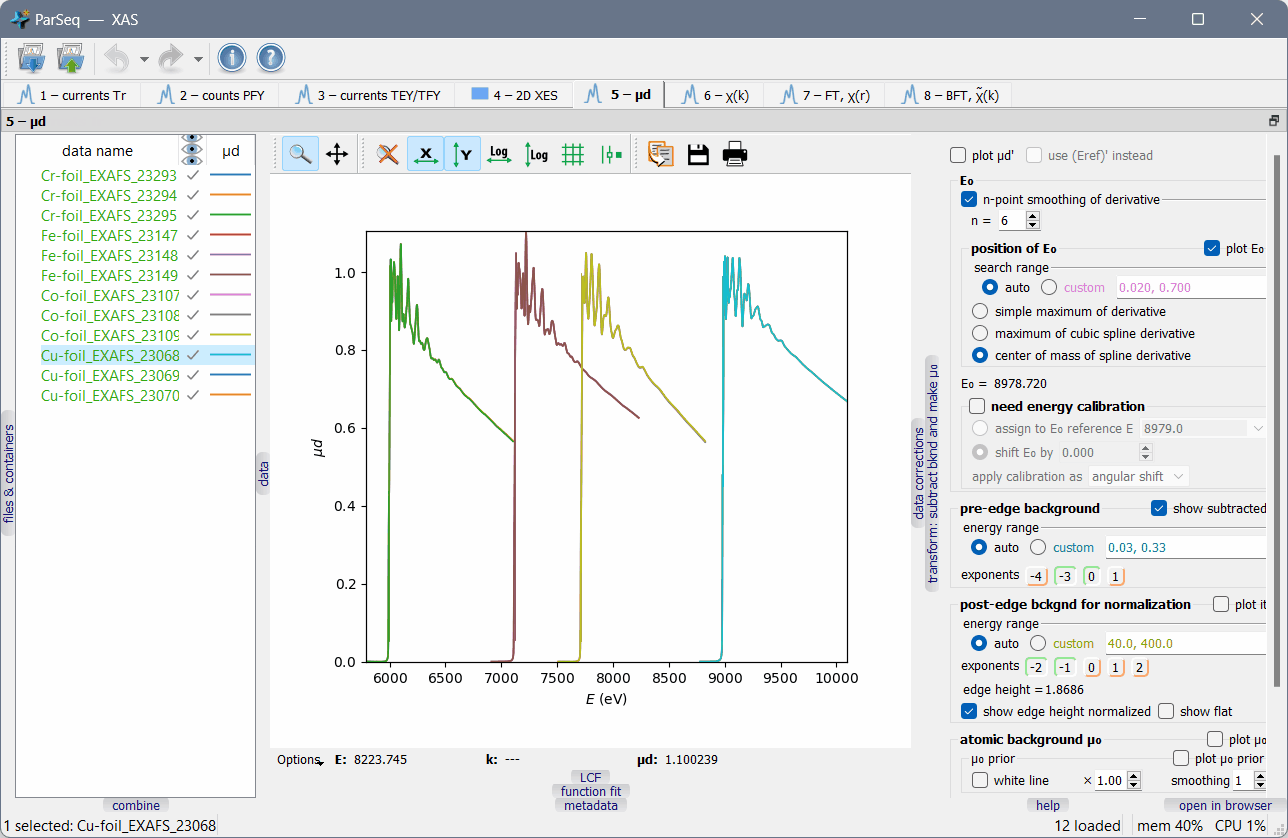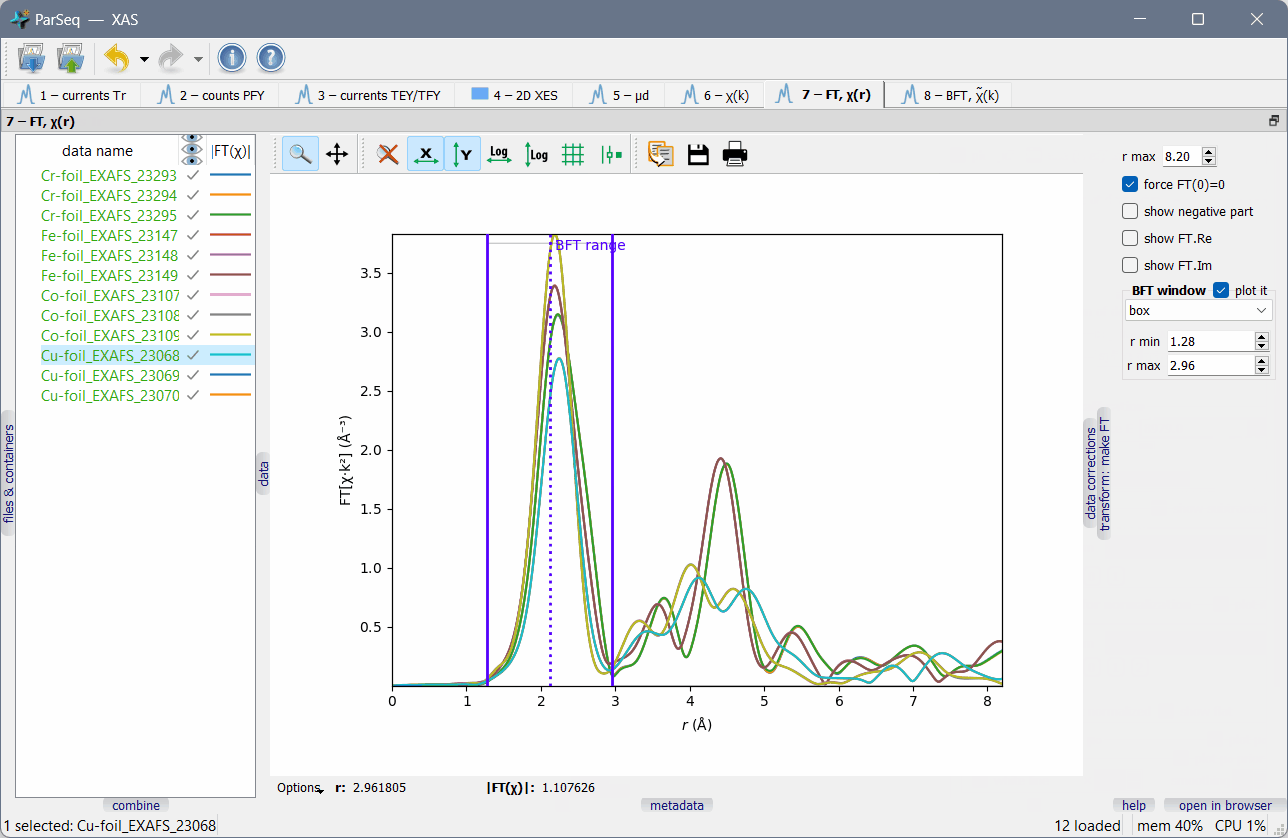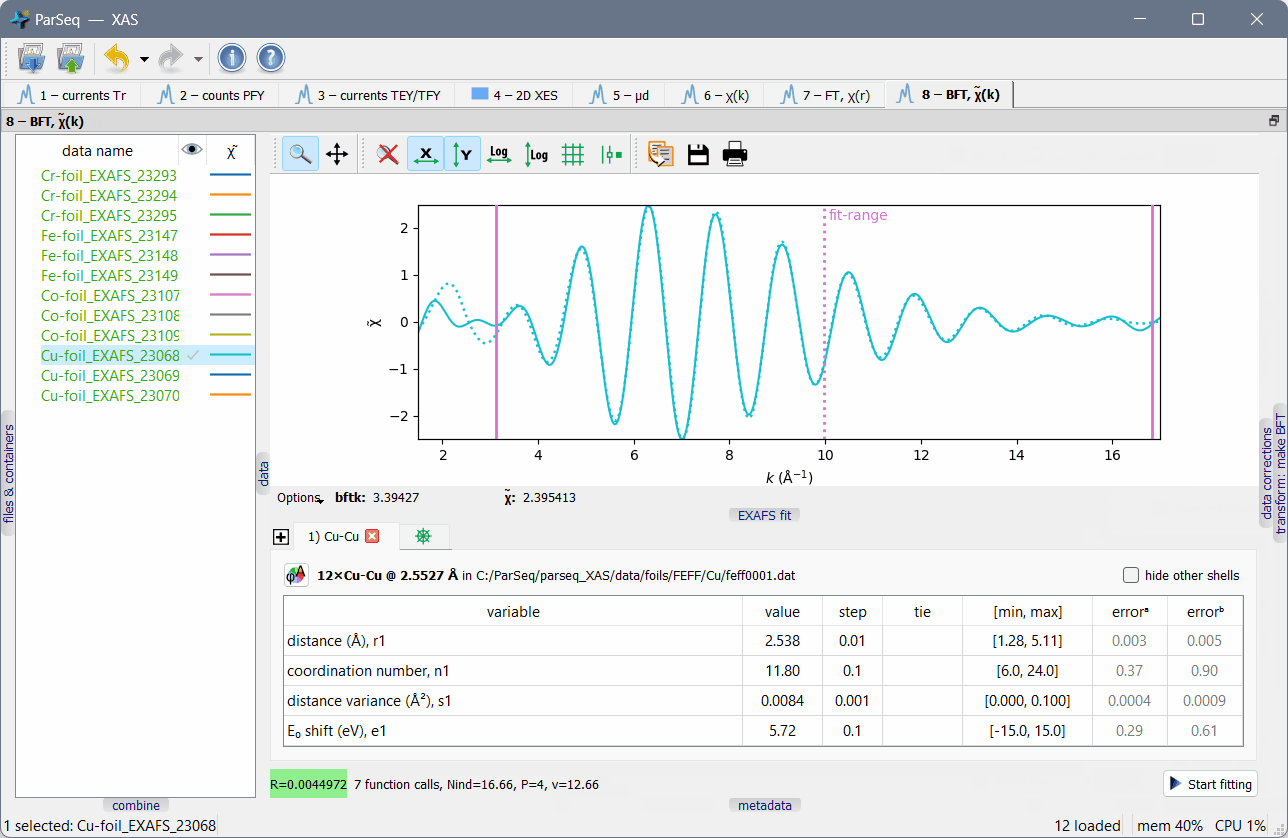
Main features¶
ParSeq allows creating analysis pipelines as lightweight Python packages.
Flexible use of screen area by detachable/dockable transformation nodes (parts of analysis pipeline).
Two ways of acting from GUI onto multiple data: (a) simultaneous work with multiply selected data and (b) copying a specific parameter or a group of parameters from active data items to later selected data items.
Undo and redo for most of treatment steps.
Entering into the analysis pipeline at any node, not only at the head of the pipeline.
Creation of cross-data combinations (e.g. averaging, PCA) and their propagation downstream the pipeline together with the parental data. The possibility of termination of the parental data at any selected downstream node.
General data correction routines for 1D data: range deletion, scaling, replacement by a spline, deletion of spikes and jump correction.
Parallel execution of data transformations with multiprocessing or multithreading (can be opted by the pipeline application).
Optional curve fitting solvers, also executed in parallel for multiple data items.
Informative error handling that provides alerts and stack traceback with the type and location of the occurred error.
Optional time profiling of the pipeline, as controlled by a command-line argument.
Export of the workflow into a project file. Export of data into various data formats with accompanied Python scripts that visualize the exported data in publication-quality plots.
ParSeq understands container files (presently only hdf5) and adds them to the system file tree as subfolders. The file tree, including hdf5 containers, is lazy loaded thus enabling big data collections.
A web viewer widget near each analysis widget displays help pages generated from the analysis widget doc strings. The help pages are built by Sphinx at the startup time.
The pipeline can be operated by the GUI or from a Python script without GUI.
Optional automatic loading of new data during a measurement time.
The mechanisms for creating nodes, transformations and curve fitting solvers, connecting them together and creating Qt widgets for the transformations and and curve fits are exemplified by separately installed analysis packages:
Running without installation¶
Get the zip files of ParSeq and a ParSeq pipeline from GitHub. Unzip their folders (parseq and e.g. parseq_XES_scan) to the same suitable directory, install dependencies and run the pipeline starter. One advantage of no installation is a single location of parseq served by any Python installation.
Running with installation¶
Install py pip:
pip install parseqand e.g.pip install parseq-XAS.Or install from unzipped GitHub archives: from folders with pyproject.toml run
python -m pip install ..
After installation, the pipeline starters are runnable from command line, e.g.
as parseq-XAS.
Launch an example¶
Run the *_start.py module of the pipeline (also available as commands if
ParSeq and the ParSeq pipeline were installed, not just unzipped). You can try
it with --help to explore the available options. An assumed usage pattern
is to load a project .pspj file from GUI or from the starting command line.
Documentation¶
The documentation is available online on Read the Docs.
Hosting and contact¶
The ParSeq project is hosted on GitHub. Please use the project’s Issues tab to get help or report an issue.
Citing ParSeq¶
Please cite ParSeq as: K Klementiev, “ParSeq: Python software for comparative data analysis pipelines”, J. Phys.: Conf. Ser. 3010 (2025) 012126; doi:10.1088/1742-6596/3010/1/012126.
User’s Guide¶
- Create analysis pipeline
- Centralized facilities
- Nodes and transformations
- Docked node widgets
- Undo and redo
- File tree views and file formats
- Metadata
- Plots
- Fits
- Cross-data combinations
- Standard data corrections
- Project saving with data export and plot script generation
- Error handling
- Command-line interface and start options
- Performance profiling
- Help system
- About dialog
- Prepare pipeline metadata and images
- Make data nodes
- Make data transformations
- Make GUI widgets
- Make fitting worker classes
- Make data pipeline
- Create test data tree
- Create pipeline starter
- Creating development versions of analysis application
- Centralized facilities
- Data nodes
- Data model
- Data transformations
- Data corrections
- Data correction widgets
- User transformation widgets
- Fitting workers
- Fit widgets
- Citing ParSeq
- The MIT License





